Presentations
Here is a list of my most recent presentations, mostly given at seminars and workshops:
-
8-Oct-2015 - Journées du GDR BIM, Institut Pasteur, Paris
The overall architecture of an RNA can be predicted with acceptable accuracy using established computational methods, such as those implemented in the MFold or the RNAFold software. This opens the door for a rational design of RNA molecules, to serve as building blocks in synthetic biology, or as novel therapeutic agents. To that purpose, one must solve the inverse folding problem, i.e. construct an RNA sequence which adopts a specific secondary structure according to established structure prediction methods. However, contrasting with the prediction task, the underlying computational structure of inverse folding is poorly understood, and existing tools either fail to return admissible sequences for some structures, or require extraordinary computational resources. Consequently, the field of RNA bioinformatics is currently craving for both practical and theoretically-sound approaches for the problem. In this talk, I will introduce the many flavors of the RNA design problem and illustrate its relevance to the objectives of modern (synthetic) biology. I will outline the strengths and shortcomings of selected approaches, and will briefly present recently contributed methods, developed with collaborators at Univ McGill (Canada), Wuhan Univ. (China) and Univ. Paris-Sud. If time allows, I will also mention results recently obtained with collaborators at Simon Fraser University, which characterize large subclasses of (un)designable secondary structures in simple energy models. These results are currently being extended to more realistic energy models, and could serve as a foundation for exact, computationally-efficient, solutions for RNA design. -
23-Jul-2015 - Benasque RNA meeting, Spain
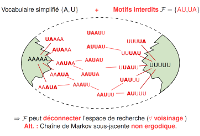
We considered the Combinatorial RNA Design problem, a minimal instance of the RNA design problem which aims at finding a sequence that admits a given target as its unique base pair maximizing structure. We provide complete characterizations for the structures that can be designed using restricted alphabets. Under a classic four-letter alphabet, we provide a complete characterization of designable structures without unpaired bases. When unpaired bases are allowed, we provide partial characterizations for classes of designable/undesignable structures, and show that the class of designable structures is closed under the stutter operation. Membership of a given structure to any of the classes can be tested in linear time and, for positive instances, a solution can be found in linear time. Finally, we consider a structure-approximating version of the problem that allows to extend bands (helices) and, assuming that the input structure avoids two motifs, we provide a linear-time algorithm that produces a designable structure with at most twice more base pairs than the input structure.
This is joint work with Jozef Hales (SFU, Canada), Jan Manuch (UBC/SFU, Canada) and Ladislav Stacho (SFU, Canada). -
6-Jun-2015 - CanaDAM'15@Saskatoon, Canada
In this talk, I emphasized how, over the past three decades, RNA biology has benefited from a continuous and fruitful cross-talk between discrete mathematicians, computer scientists and biochemists. At the center of this conversation lies the concept of dynamic-programming, an algorithmic design technique which solves a combinatorial optimization problem efficiently by taking advantage of a well-chosen decomposition of its search space. Extensions and optimized instances of this technique now allow to address, at a genomic scale, multiple questions related to the analysis of the Boltzmann ensemble and the sequence-structure(-function) relationship. These developments also raise well-defined open questions, motivating further studies of the underlying discrete structures. -
12-Jun-2014 - Scientific Café of the French consulate@Alliance Française@Vancouver, Canada
Scientific outreach talk during an event hosted by Alliance Française. As usual, this deals with RNA and dynamic programming, but using more colors and less equations than usual! -
10-Jun-2014 - Discrete Maths Seminar@SFU, Canada
RiboNucleicAcids (RNAs) are ubiquitous and versatile molecules encoded in our DNA, and the object of much interest in modern biology. The function of an RNA in biological systems largely depends of its structure, which can be mathematically abstracted as an arc-annotated sequence with possible additional constraints (crossing/non-crossing, single/bounded list/arbitrary partners per position). Accordingly, one of the foremost problem in RNA computational biology is the search, in genomes of sequenced organisms, for genes encoding -- previously undetected -- RNAs.
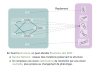
In this talk, we consider RNA structure as a dependency graph, which connects positions in the structure that have to be simultaneously considered in order to correctly optimize the distance. We then introduce a dynamic-programming algorithm which computes the best sequence/structure alignment, starting from a tree-decomposition of the dependency graph. The complexity of the algorithm is linear on the structure length, and exponential on the width of its tree-decomposition. We propose some decomposition heuristics for computing the tree-decomposition of the dependency graphs. The two-steps algorithm (structure->tree-decomposition->alignment) specializes, on each of the previously investigated classes, into algorithms having best-known complexities.
This is joint work with Philippe Rinaudo (Univ. Paris-Sud), Alain Denise (Univ. Paris-Sud) and Dominique Barth (Univ. Versailles-St Quentin) -
12-May-2014 - Lecture@EPIT'14 - Oléron Island, France
This joint lecture (in French!) with Alain Denise illustrates some basic and advanced instances of dynamic programming applied to RNA computational biology. -
15-Apr-2014 - PARC@SFU, Canada
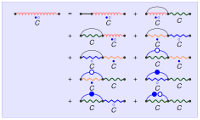
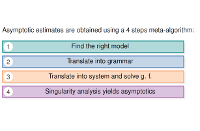
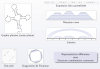
In this talk, we recall the basic tools of singularity analysis, and show how they can be used, within a simple four-steps program, to study the asymptotic properties of RNA secondary structures. The whole methodology specializes the symbolic method introduced by Flajolet et al, and allows for multiple extensions, such as the incorporation of weights to model Boltzmann-like distributions. We illustrate this unified framework by deriving various quantities, such as the average distance between the extremities of an RNA, the distribution of occurrences of motifs in a random structure, the average complexity of RNA sampling algorithms, or the asymptotic growth of RNA shapes, some abstract representation of RNA structure. Some of these results, in comparison with experimental data, reveal discrepancies which question the representativity of "pure" combinatorial models (eg homopolymer hypothesis). -
17-Mar-2014 - ALEA'14@CIRM, Marseille
Overview at ALEA'14 of recent collaborative work relying on a random generation (aka global sampling) paradigm to tackle the RNA design problem. Joint work with multiple collaborators at UCSC (USA), Paris XI (France), Wuhan univ. (Chine), LIGM/Paris-Est (France), MIT (USA) and McGill (Canada). -
23-Sep-2013 - ACM-BCB'13@Washington, USA
Presentation of an extension of our global sampling method for RNA design, which allows for rich sets of constraints while remaining in linear time. Joint work with Y. Zhou (UCSC, USA), A. Denise(LRI/IGM, Paris XI), Y. Zhang (Wuhan Univ., Chine), S. Vialette (LIGM, Paris-Est) and J. Waldispühl (McGill, Canada). -
15-Feb-2013 - TBI WinterSeminar@Bled, Slovenia
Invited talk at the Winter Seminar of the Theoretical Biochemsitry Institute (Vienna, home of the infamous Vienna RNA Package!), describing a recently contributed parameterized-complexity algorithm based on tree-decomposition. This work was contributed jointly with P. Rinaudo, A. Denise et D. Barth (Slidesshamefully rippedadapted from P. Rinaudo's PhD defense). -
28-Oct-2012 - Researcher' night@Polytechnique
Popular science event, organized at Ecole Polytechnique. Following a long-time collaboration with the communication of Inria Saclay, we designed and presented several activities related to RNA algorithmics. In addition to the slides (alas, in French :( ), I coded two small pieces of software for the occasion: a minimal implementation of a Nussinov-style RNA folding algorithm [Jar], and an even more minimalist RNA design studio [Jar] (Eterna mockup). -
4-Jul-2012 - CPM'12@Helsinki, Finland
Predicting the folding of an RNA sequence, while allowing general pseudoknots (PK), consists in finding a minimal free-energy matching of its n positions. Assuming independently contributing base-pairs, the problem can be solved in O(n3)-time using a variant of the maximal weighted matching. By contrast, the problem was previously proven NP-Hard in the more realistic nearest-neighbor energy model.
With Rolf Backofen (Uni Freiburg, Germany) and Saad Sheikh (University of Florida, USA), we considered an intermediate model, called the stacking-pairs energy model. We extended a result by Lyngso, showing that RNA folding with PK is NP-Hard within a large class of parametrization for the model. We also showed the general approximability of the problem, by giving a basic O(n3) algorithm that achieves at least a 5-approximation for any parametrization of the stacking model. This contrasts nicely with the nearest-neighbor version of the problem, which we proved cannot be approximated within any positive ratio, unless P=NP. -
18-Jun-2012 - AOFA'12@Montreal, Canada
Presentation of a study, carried out in a collaboration with Jérémie Du Boisberranger and Danièle Gardy (Prism, Université de Versailles St Quentin, France), of a non-uniform instance of the coupon collector problem. Inspired by bioinformatics, we consider weighted languages and give a general (yet not universal) theorem for the waiting time of the full collection. -
12-Jun-2012 - IGM RNA retreat@Seillac, France
A joint tutorial with Alain Denise (LRI/IGM, Paris XI, France) on currently available methods for RNA structure prediction. -
6-Apr-2012 - Unithé ou Café, INRIA Saclay, France
Science for the masses! :) Foundations of RNA folding algorithms explained as a folkloric dance (in French). -
6-Mar-2012 - VIZBI'12, EMBL Heidelberg,Germany
Invited talk giving a broad overview of the existing visualization techniques/methods/tools and many challenges faced by the RNA community. -
5-Mar-2012 - VIZBI'12, EMBL Heidelberg,Germany
-
5-Sep-2011 - WABI'11, Saarbrücken, Germany
Presentation at WABI'11 (Saarbrücken, Germany) of a joint work with C. Saule (LRI/IRIC) using hypergraphs as a formalism to describe and develop pseudoknotted algorithms. -
25-May-2011 - 12th Maths Culture and Games exhibition, Paris France
Participation on the behalf of INRIA at a popular science event, the 12th Maths Culture and Games exhibition, located in Paris. Our presentation was dedicated to RNA folding (Check out our cool lightweight implementation of the Nussinov algorithm, used in the event to display the superiority of elegant algorithmic solutions over brute force approaches) and co-animated with M. Regnier, F. Lou, P. Rinaudo, V.-D. Tran, and A. Guilhot-Gaudeffroy (INRIA AMIB). -
25-Apr-2011 - IWBC/McGill@Barbados'11
Presentation at the "1st Integrative Workshop on Biomolecular Complexes" of combinatorial techniques underlying statistical sampling methods and application to the exploration of the mutational landscape. -
31-Mar-2011 - RECOMB'11@Vancouver, CA
Présentation of a novel adaptive sampling algorithm for accessing regions of the mutational landscape. -
24-Feb-2011 - Sequence Processing'11@Dagstuhl
Hypergraph framework and dynamic programming for the RNA folding problem with pseudoknots. Presentation of natural extensions toward Ensemble applications. -
18-Mar-2011 - MMC seminar@Polytechnique
Overview talk on the thermodynamical aspects of RNA folding throughout evolution, given at the Models for Cell Metabolism multidisciplinary seminar at Ecole Polytechnique. -
14-Nov-2010 - ROC meeting@Strasbourg
Presentation of VARNA's most recent extensions and of possible ways to represent alignements information visually and/or interactively. -
3-Sep-2010 - Talk@GASCOM'10 (Montréal, Canada)
Presentation, given at UQAM (Montréal, Canada), of a joint effort with Danièle Gardy (UVSQ) to analyze the level of redundancy of a random generation of samples within a weighted (aka Boltzmann) distribution. -
2-Sep-2010 - Computational Models for RNA@McGill (Montréal, Canada)
Talk at the McGill center for bioinformatics (Montreal, Canada) within a "Computational Models for RNA structures" serie, presenting ongoing works on an Hypergraph analogy for RNA dynamic programming. -
22-Mar-2010 - LACL seminar (Paris XII/Créteil)
A slightly extended version of ALEA's talk. -
22-Mar-2010 - ALEA workshop (Luminy)
Combinatorial re-interpretation of RNA folding algorithms. -
25-Feb-2010 - ARENA workshop (Boussens)
Presentation (in French) of the thermodynamic and algorithmic principles used for the equilibrium prediction of RNA folding. -
25-Feb-2010 - ARENA workshop (Boussens)
Short presentation (in French) of most recent extensions to our VARNA software: Session save/load, region annotations... -
15-Dec-2009 - OCAD seminar (LIPN/Paris XIII-France)
Presentation of some results on combinatorial problems arising in the study of RNA folding. This talk puts a special emphasis on a combinatorial interpretation of the conformational space, which provides a unifying view over MFE folding, partition function and statistical sampling algorithms. -
27-Nov-2009 - Bioinformatics group seminar (Freiburg university-Germany)
Short introduction of the so-called symbolic method and its extension (through singularity analysis) to the automatic extraction of asymptotic behaviors for simples combinatorial classes (Regular and context-free languages). Application to the analysis of RNA shapes, a coarse-grain representation of RNA secondary structure. -
20-Mar-2009 - ALEA workshop (Luminy)
Principles and preliminary results of multidimensional random generation. Collaborative work with O. Bodini (LIPN, Paris XIII). -
23-Jun-2008 - GASCOM'08 Conférence (Bibbiena-Italia)
Presentation of a new random generation algorithm. This algorithm is non-redundant, in the sense that it avoids any previously generated word while remaining unbiased (e.g. uniform) on the remaining set of objects.